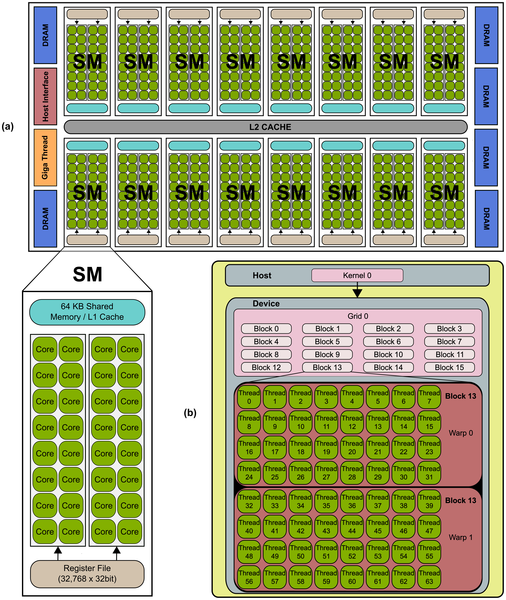
Current processors are endowed with many simpler processors, having a tremendous potential in terms of peak performance. Moreover, emergent platforms such as Graphics Processing Units (GPUs), the Field Programmable Gate Array (FPGAs), Accelerating Processing Units (APUs), etc. have been consolidated for developing scientific applications in different areas including bioinformatics, finance, seismic Processing, fluid dynamics, etc. However, it is not a trivial task to take advantage of the potential performance that these platforms provide to the scientific community. In this research line, we develop scientific application from different fields such as linear algebra, system biology, natural computing, image processing, etc. On these emergent platforms, providing insight into the peculiarities of their programming models and architectures. Currently, we are researching in applying those models to challenging problems such as those coming from Bioinformatics.
Some previous group results
Q. Zhang, G.D. Guerrero, J.M. Cecilia, J.M. García, J. Wang, T. Hou, H. Pérez-Sánchez, (2013) “A enhanced GPU based Conformational Entropy Calculation Method”, Journal of Chemical Information and Modeling, (in press).
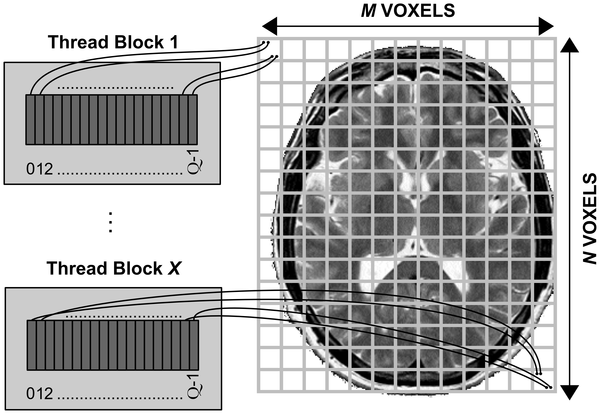
M. Hernández, G. Guerrero, J. M. Cecilia, José M. García, A. Inuggi, et al. Accelerating Fibre Orientation Estimation from Diffusion Weighted Magnetic Resonance Imaging Using GPUs.. (2013) PLoS ONE 8(4): e61892. doi:10.1371/journal.pone.0061892
I. Sánchez-Linares, H. Pérez-Sánchez, J.M. Cecilia, J.M. García, (2012) “High-Throughput parallel blind Virtual Screening using BINDSURF”,BMC Bioinformatics, 13(Suppl 14):S13 (Highly accessed).
José M. Cecilia, José M. García, Manuel Ujaldón, Andy Nisbet and Martyn Amos. Parallelization Strategies for Ant Colony Optimisation on GPUs. In 14th Int. Workshop on Nature Inspired Distributed Computing -NIDISC11- (in conjunction with IPDPS 2011), Anchorage (Alaska), May 2011.